RNA isolation from plant tissues
The plant material surveyed in this project was kindly provided by the laboratories reported in the Species List. Three different RNA isolation protocols/reagents, capable of retaining the small RNA fraction, were selected and optimized to extract high quality total RNA from all the samples in the panel:
1) Plant RNA Purification Reagent® (PRPR), Invitrogen
2) TRIZOL® (Invitrogen) / TRIREAGENT® and TRIREAGENT LS® / (Molecular Research Center)
3) Adapted from Bugos et al. (BioTechniques 1995, vol. 19:731-737).
Quality of total RNA was evaluated by agarose gel electrophoresis, micro-electrophoresis (Agilent 2100 Bioanalyzer®, Ambion) and spectrophotometry (Nanodrop ND-1000®).
Technical details on these procedures and further insights into the processing of special tissues are discussed in Accerbi et al., 2008 (“Total RNA isolation for small RNA analysis” a chapter in the forthcoming book entitled “Plant microRNA protocols”, in the Methods in Molecular Biology series).
Small RNA libraries
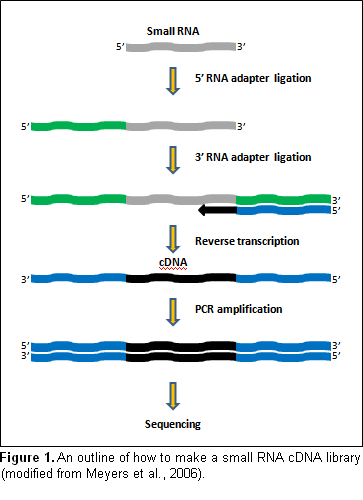
RNA samples were submitted to Illumina (Hayward, CA, http://www.illumina.com) for small RNA library construction using approaches described in Lu et al. (2007) Methods 43:110-117, with minor modifications. The method begins with the precipitation of LMW RNA from total RNA and the purification of small RNAs by polyacrylamide gel-based size fractionation. After ligation of RNA adapters to small RNAs, a low number of PCR cycles is used to obtain sufficient quantities of template (Figure 1). The small RNA libraries are then sequenced with Solexa’s SBS technology, with sequencing performed by Illumina.
Sequencing
In this project, we are using SBS sequencing, as it provides full-length small RNA sequences at considerable depth (approximately 1 to 3+ million reads per library). Information about the SBS sequencing method is available from the company website (http://www.illumina.com). For a general overview of commonly used small RNA cloning and sequencing methods, see Meyers et al. (2006) Curr. Op. in Biotech., 17:139-146.